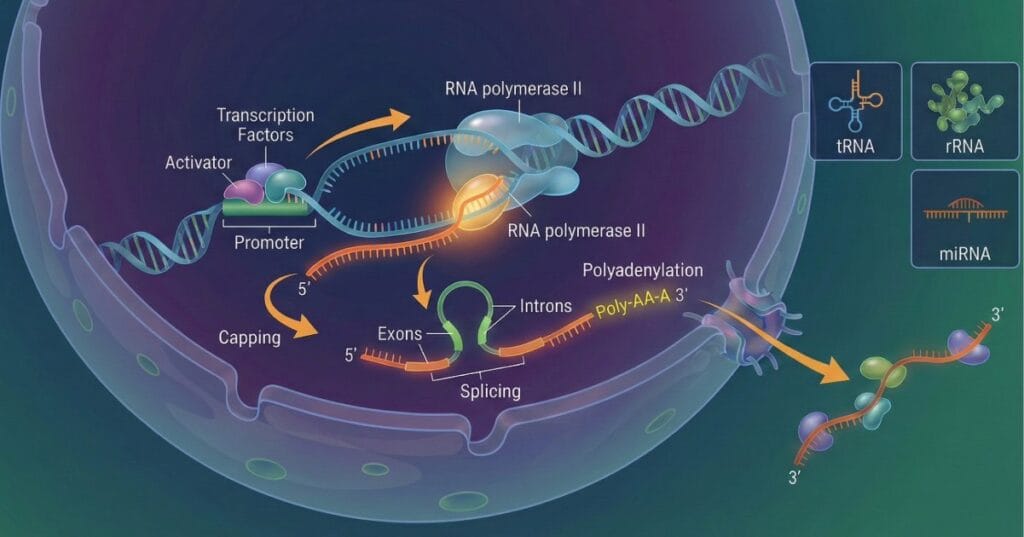
COMPETITIVE EXAM MCQs SERIES of LIFE SCIENCES for CSIR-UGC NET/JRF, SLET, GATE, and other entrance tests: FUNDAMENTAL PROCESSES – RNA Synthesis and Processing.
Syllabus Outline
- Transcription factors and machinery.
- Formation of the initiation complex.
- Transcription activator and repressor.
- RNA polymerases, capping, elongation, and termination.
- RNA processing, RNA editing, splicing, and polyadenylation.
- Structure and function of different types of RNA.
- RNA transport.
This quiz contains concept-based, most frequently asked 25 MCQs of “FUNDAMENTAL PROCESSES – RNA Synthesis and Processing”. Each question has a single correct/most appropriate answer.
*****
1. In eukaryotic RNA Polymerase II, the transition from transcription initiation to elongation is chemically signalled by the phosphorylation of which residue in the C-terminal domain heptad repeat?
A) Serine 2
B) Serine 5
C) Serine 7
D) Tyrosine 1
2. Which of the following eukaryotic RNA polymerases is responsible for the synthesis of the 5S ribosomal RNA?
A) RNA Pol I
B) RNA Pol II
C) RNA Pol III
D) Mitochondrial RNA Pol
3. The 5′ capping of eukaryotic pre-mRNA is a co-transcriptional process. Which of the following enzymatic activities is NOT required for the formation of the m7G cap?
A) RNA Triphosphatase
B) Guanylyltransferase
C) Methyltransferase
D) Adenylyltransferase
4. RNA Polymerase III promoters are unique among eukaryotic polymerases because they often contain regulatory elements that are:
A) Located 1 kb upstream of the start site.
B) Located within the transcribed region.
C) Recognised directly by the TATA-binding protein without TAFs.
D) Sensitive to extremely low concentrations of alpha-amanitin.
5. The first transesterification reaction in nuclear pre-mRNA splicing involves the nucleophilic attack of which chemical group on the 5′ splice site?
A) The 3′-OH of the 5′ exon.
B) The 2′-OH of the branch point Adenosine.
C) The 5′-P of the branch point Adenosine.
D) The 2′-OH of the 5′ splice site Guanosine.
6. RNA editing by the Adenosine Deaminase Acting on RNA family of enzymes results in the conversion of Adenosine to Inosine. During translation, Inosine is read as which nucleotide?
A) Uracil
B) Cytosine
C) Guanosine
D) Thymidine
7. Which modification of the C-terminal domain of RNA Pol II is specifically associated with the recruitment of the 3′ end processing and polyadenylation machinery?
A) Phosphorylation of Ser5
B) Phosphorylation of Ser2
C) Acetylation of Lysine residues
D) Methylation of Arginine residues
8. What is the consensus sequence for the cleavage of the 3′ end of the nascent pre-mRNA?
A) GGGCCC
B) AAUAAA
C) TATAAT
D) AGGTGA
9. During the RNA transport, the Exon Junction Complex is typically deposited on the mRNA at which position?
A) 100 nucleotides downstream of the exon-exon junction.
B) Exactly at the exon-exon junction.
C) 20-24 nucleotides upstream of the exon-exon junction.
D) Attached to the poly(A) tail.
10. Which of the following statements regarding RNA Polymerase I is TRUE?
A) It transcribes tRNAs and 5S rRNA.
B) It is highly sensitive to alpha-amanitin.
C) It transcribes a single large precursor RNA in the nucleolus.
D) It requires the TFIIB factor for promoter recognition.
11. In mitochondrial RNA editing in Trypanosomes, the insertion of Uracil residues into the mRNA is facilitated by which enzyme?
A) RNA Polymerase II
B) Terminal Uridylyl Transferase
C) Trypanosomase
D) Poly(U) Polymerase
12. The “Wobble” base in a tRNA anticodon allows it to recognise multiple codons. If the 5′ base of the anticodon is Inosine (I), it can pair with which of the following bases at the 3′ position of the codon?
A) A and G only
B) U, C, and A
C) U and C only
D) G only
13. Self-splicing Group I introns utilise an external cofactor for the first transesterification reaction. What is this cofactor?
A) ATP
B) Adenosine
C) Magnesium ions
D) Guanosine
14. RNA Polymerase II termination at the 3′ end of a gene is coupled with the cleavage of the transcript. Which model suggests that an exonuclease “catches up” to the polymerase to trigger termination?
A) Allosteric model
B) Torpedo model
C) Recruitment model
D) Loop model
15. Which of the following situations is most likely to trigger nonsense-mediated mRNA decay in eukaryotic cells?
A) Translation termination at a stop codon located in the last exon of an mRNA
B) Translation termination at a stop codon located within 20 nucleotides of the poly(A) tail
C) Presence of a stop codon within the 5′ untranslated region
D) Translation termination at a stop codon located upstream of a downstream exon junction complex
16. The Sex-lethal gene in Drosophila regulates sex-specific splicing. In females, the Sex-lethal protein acts as a:
A) Transcriptional activator of the transformer
B) Component of the m7G capping complex.
C) Kinase that phosphorylates the TRA protein.
D) Splicing repressor that binds to the transformer pre-mRNA.
17. RNA polymerase II contains a bridge helix that is essential for enzyme translocation during transcription elongation. The toxin α-amanitin inhibits RNA polymerase II by binding near this helix. What is the primary molecular consequence of this interaction?
A) It blocks the entry of nucleoside triphosphates into the active site.
B) It induces immediate cleavage of the 3′ end of the nascent RNA.
C) It causes dissociation of the pre-initiation complex.
D) It inhibits the physical translocation of RNA polymerase II along the DNA template.
18. The process of trans-splicing, in which exons from two different primary RNA transcripts are joined together to form a mature mRNA, is a characteristic feature of gene expression in which organism?
A) Escherichia coli
B) Saccharomyces cerevisiae
C) Trypanosoma brucei
D) Drosophila melanogaster
19. Which modification of the DNA template is most commonly associated with transcriptional silencing in eukaryotic cells?
A) Histone acetylation
B) DNA methylation at CpG islands
C) Phosphorylation of histone H3 at serine 10
D) Trimethylation of histone H3 at lysine 4
20. The following statements are true about the processing of eukaryotic pre-mRNA:
I – The 5′ capping enzyme is recruited to the RNA Pol II C-terminal domain when it is phosphorylated at Serine 5.
II – Splicing involves two sequential transesterification reactions that result in the ligation of exons.
III – The poly(A) tail is added to the 3′ end of mRNA by RNA Polymerase II during termination.
IV – Splicing can only occur after the entire pre-mRNA transcript has been released from the DNA.
A) I only
B) II only
C) I and II
D) I, II, III and IV
21. Which of the following statements are true about RNA editing:
I – A-to-I editing is catalysed by the Adenosine Deaminases acting on RNA enzyme family.
II – C-to-U editing in the ApoB gene creates a premature stop codon in liver cells.
III – Guide RNAs are required for U-insertion editing in Trypanosomes.
IV – RNA editing can alter the amino acid sequence of a protein without changing the genomic DNA.
A) I, III and IV
B) II, III and IV
C) I, II and III
D) I and IV only
22. Assertion (A): The mRNA produced in eukaryotes is generally monocistronic.
Reason (R): Eukaryotic ribosomes initiate translation by recognising the 5′ cap and scanning for the first AUG codon, which prevents internal initiation on polycistronic messages.
A) Both (A) and (R) are true, and (R) is the correct explanation of (A).
B) Both (A) and (R) are true, but (R) is not the correct explanation of (A).
C) (A) is true, but (R) is false.
D) (A) is false, but (R) is true.
23. Assertion (A): Splicing of pre-mRNA is an energy-generating process.
Reason (R): Four ATPs are generated during the two transesterification reactions for intron removal and exon ligation.
A) Both (A) and (R) are true, and (R) is the correct explanation of (A).
B) Both (A) and (R) are true, but (R) is not the correct explanation of (A).
C) (A) is true, but (R) is false.
D) Both (A) and (R) are false.
24. If a eukaryotic cell is incubated with radioactive 32P labelled at the beta-phosphate position of ATP, where will the radioactivity appear in the mature mRNA?
A) In the phosphodiester backbone of the RNA chain.
B) Only in the 5′ cap structure.
C) It will not appear in the mRNA as beta phosphate is released during polymerisation.
D) Uniformly distributed throughout the poly(A) tail.
25. Which of the following would be the most likely cellular phenotype if the Exportin-5 gene is mutated?
A) Failure to export all mRNAs.
B) Failure to export tRNAs.
C) Accumulation of pre-miRNAs in the nucleus.
D) Ribosomal subunits cannot enter the cytoplasm.
*****
Previous: DNA Replicon and Recombination
Next: Protein Synthesis and Processing
References
- Nelson, David L. & Cox, Michael M. (2021). Lehninger Principles of Biochemistry, W. H. Freeman, 8th Edition.
- Alberts, B., Johnson, A., Lewis, J., Morgan, D., Raff, M., Roberts, K., & Walter, P. (2014). Molecular Biology of the Cell, Garland Science, 4th Edition.
- Geoffrey Cooper and Kenneth Adams (2022). The Cell: A Molecular Approach, Oxford University Press, 9th Edition
- Kuby, J., Kindt, T. J., Osborne, B. A., & Goldsby, R. A. (2019). Kuby Immunology, W. H. Freeman, 8th Edition.
- Gupta, P.K. (2022). Cell and Molecular Biology, Rastogi Publications, 5th Edition.
🔗 Explore More MCQs: